Ambitious survey of human diversity yields millions of undiscovered genetic variants
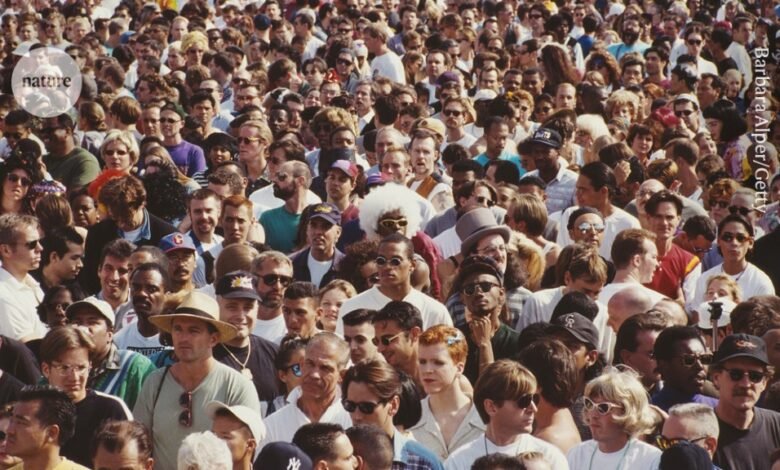
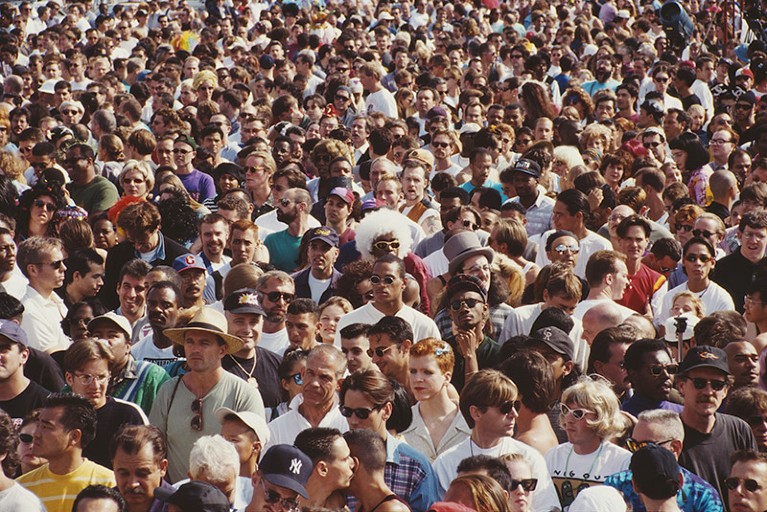


The All of Us programme aims to recruit one million people from ethnic and socio-economic groups that are typically under-represented in biomedical studies.Credit: Barbara Alper/Getty
A massive US programme that aims to improve health care by focusing on the genomes and health profiles of historically underrepresented groups has begun to yield results.
Analyses of up to 245,000 genomes gathered by the All of Us programme, run by the US National Institutes of Health in Bethesda, Maryland, have uncovered more than 275 million new genetic markers, nearly 150 of which might contribute to type 2 diabetes. The work has also identified gaps in genetics research on non-white populations. The findings were published on 19 February in a package of papers in Nature1,2, Communications Biology3 and Nature Medicine4.
They are a “nice distillation of the All of Us resource — what it is and what it can do”, says Michael Inouye, a computational genomicist at the University of Cambridge, UK. “This is going to be the go-to data set” for genetics researchers who want to know whether their findings are generalizable to a broad population or apply to only a limited one, he adds.
Bridging the gap
Researchers have long acknowledged the lack of diversity in the genomes available for them to study, says Jibril Hirbo, a geneticist at Vanderbilt University Medical Center in Nashville, Tennessee, who studies the genetics of health disparities. One study5 that looked at data gathered up until January 2019 found that 78% of people in most large-scale genomic studies of disease were of European descent. This has exacerbated existing health disparities, particularly for non-white individuals, Hirbo says. When researchers choose genetic or molecular targets for new medicines or create models to predict who is at risk of developing a disease, they tend to make decisions on the basis of non-diverse data because that’s all that has been available.
Facing up to injustice in genome science
The All of Us programme, which has received over US$3.1 billion to date and plans to assemble detailed health profiles for one million people in the United States by the end of 2026, aims to bridge that gap, says Andrea Ramirez, the programme’s chief data officer. It began enrolling people in 2018, and released its first tranche of data — about 100,000 whole genomes — in 2022. By April 2023, it had enrolled 413,000 anonymized participants, 46% of whom belong to a minority racial or ethnic group, and had shared nearly 250,000 genomes. By comparison, the world’s largest whole-genome data set, the UK Biobank, has so far released about half a million genomes, around 88% of which are from white people.
The All of Us data set is “a huge resource, particularly of African American, Hispanic and Latin American genomes, that’s massively missing from the vast majority of large-scale biobank resources and genomics consortia”, says Alicia Martin, a population geneticist at Massachusetts General Hospital in Boston.
In addition to the genomes, the database includes some participants’ survey responses, electronic health records and data from wearable devices, such as Fitbits, that report people’s activity, “making this one of the most powerful resources of genomic data”, Martin says.
An urgent need
A study in Nature on type 2 diabetes2 is an example of the power of using a database that includes diverse genomes, Ramirez says. The condition, which affects about one in ten people in the United States, can be caused by many distinct biological mechanisms involving various genes. The researchers analysed genetic information from several databases, including All of Us, for a total of more than 2.5 million people; nearly 40% of the data came from individuals not of European ancestry. The team found 611 genetic markers that might drive the development and progression of the disease, 145 of which have never been reported before. These findings could be used to develop “genetically informed diabetes care”, the authors write.
World’s biggest set of human genome sequences opens to scientists
In another of the studies3, researchers used All of Us data to examine pathogenic variants — that is, genetic differences that increase a person’s risk of developing a particular disease. They found that, among the genomes of people with European ancestry, 2.3% had a pathogenic variant. Among genomes from people with African ancestry, however, this fell to 1.6%.
Study co-author Eric Venner, a computational geneticist at Baylor College of Medicine in Houston, Texas, cautions that there should be no biological reason for the differences. He says that the disparity is probably the result of more research having been conducted on people of European ancestry; we simply know more about which mutations in this population lead to disease. In fact, the researchers found more variants of unknown risk in the genomes of people with non-European ancestry than in those with European ancestry, he adds. This underscores the urgent need to study non-European genomes in more detail, Venner says.
Updating models
Gathering and using more genomic and health data from diverse populations will be especially important for generating more accurate ‘polygenic risk scores’. These provide a picture of a person’s risk of developing a disease as a result of their genetics.
US tailored-medicine project aims for ethnic balance
To calculate a score for a particular disease, researchers develop an algorithm that is trained on thousands of genomes from people who either do or don’t have the disease. A person’s own score can then be calculated by feeding their genetic data into the algorithm.
Previous research6 has shown that the scores, which might soon be used in the clinic for personalized health care, tend to be less accurate for minority populations than for majority ones. In one of the current papers4, researchers used the more-inclusive All of Us data to improve the landscape: they calibrated and validated scores for 23 conditions and recommended 10 to be prioritized for use in the clinic, for conditions including coronary heart disease and diabetes. Martin applauds these efforts, but she hopes that future studies address how physicians and others in the clinic interpret these scores, and whether the scores can improve a person’s health in the long term because of the treatment decisions they elicit.
The All of Us programme plans to release a tranche of data every year, representing new enrolees and genomes, including one later in 2024, Ramirez says. It’s excellent that diverse data are coming in, Hirbo says, adding that he would like to see existing algorithms that were trained mainly on the genomes of people of European ancestry updated soon. “The models are still way behind,” he says.
Source link